Comparatives Outcomes Study of Patients Hospitalized with Diabetes and Myocardial Infarction: EHR Data Interrogation Among Hospital Categories
by Priscilla O. Okunji, Ph.D.,
Howard University, College of Nursing and Allied Health Sciences
Nawar Shara, Ph.D., MedStar Health Research Institute
John Kwagyn, Ph.D., Howard University, College of Medicine
Ian Brooks, Ph.D., MedStar Health Research Institute
Thomas Mellman, MD, Howard University, College of Medicine

ABSTRACT
Electronic health records (EHRs) provide vast opportunities to improve patient care. Many challenges still exist and may hinder efforts for multi-center collaborations and comparative studies. There have been concerns regarding some challenges such as systems interoperability, obstructive data collection, and uncertain generalizability of the results. However, it is important to note that leveraging EHRs to counterbalance these trends is an area of intense interest. To inform this issue, we sought to review these issues using recent articles to inform the review. We worked with data extracted from urban hospitals categorized into A and B groups to answer the study research questions.
Methods: Using ICD-9 codes for diabetes (DM) (25000) and myocardial infarction (MI) (41000), data were extracted from urban hospitals. The data were then cleaned, merged using common fields, and analyzed. ICD-9 was used instead of ICD-10 for the data extraction because the hospital category with the ICD-9 extracted data was the legacy hospital that provided the original data and variables which made the grant award and this project possible.
Results: The demographic distribution showed that males (59.4%) were admitted more for the diabetes category than females (40.6%). In contrast, females were admitted more for MI only or DM + MI. It is also important to note that younger individuals were admitted more for the diabetes category among hospital category A inpatients when compared to category B. Age, gender and race distribution were statistically significant in the diabetes only category with a p-value (<0.001) when compared to the MI only category in age (0.005) and DM + MI category in age (0.027) and race (<0.000). For the clinical measures, Diastolic blood pressure (DBP), and High Density Lipoprotein (HDL) were statistically significant at p <0.001 for diabetes only. Overall, there was a significantly lower risk of death for hospital category A (combined) patients compared to hospital category B patients.
Conclusion: Comparative studies as preliminary studies through EHR interrogation is the future. This project has confirmed that similar studies should be encouraged and may lead to preventive health education that may ultimately prevent higher mortality rate in certain populations. This project is a proof of concept approach to how data from different EHR platforms can be used to conduct a comparative study by a direct hospital EHR interrogation, without additional time needed in bedside data collection or purchase of already collected datasets.
Keywords: Data, Hospitals, Interoperability, Electronic Health Records, Diabetes, Myocardial Infarction
Background
The endeavor to establish Electronic health records (EHR) for all Americans by 2014 was launched by President George Bush Jr. in 2004. The Clinical Decision Support aspect of EHRs deliver queues, called “just in time”, to assist healthcare providers, nurses and patients to take advantage of the use of health information systems and contributes to the reduction of cost, increase in patient provider satisfaction and improvement of efficiency. The National Electronic Disease Surveillance System (NEDSS) also has a significant role in these efforts. It provides support and development of surveillance systems that are integrated to transmit health information, securely and efficiently over the internet. This in turn allows for rapid identification of natural disease outbreaks or bioterrorism. There are several different simultaneous movements to standardize information technology and its terms. The Unified Medical Language System (UMLS) attempts to unify this process. It addresses three different but interrelated types of information. The first is the Metra thesaurus which is simply a large vocabulary database. The second is the semantic network that provides a unified systematic category to the large vocabulary database. The last is the SPECIALIST Lexicon and Lexical Program, comprised of biomedical terms. The plan for UMLS is to link EHRs with biomedical literature in the future.
EHRs were initially adopted as the primary data source for various observational, epidemiological or comparative effectiveness studies. The advancement of EHR utilization for randomized clinical trials may have opened the door for study feasibility, facilitated patient recruitment, and help to streamline data collection at baseline and follow-up study points. It is worth noting that EHR data mining presents many challenges with data security and privacy, interoperability of disparate systems and infrastructure maintenance for repeat use of high-quality data in clinical research. EHRs house structured data (clinical codes for problems; diagnoses; treatment; and management) as well as unstructured data (free text; images). EHR re-use data are mainly research focused on the structured part of the EHR, thus clinical codes are used to extract meaningful information from raw EHR data. Clinical code “represent a single clinical concept such as a diagnosis, a procedure, an observation or a medication” (Williams, Kontopantelis, Buchan, & Peek, 2017, p.1). After construction, the set is then used to query and extract data from an EHR database, for use in future analysis. The sets of codes, referred to as ‘‘code lists”, ‘‘clinical code lists”, ‘‘code sets” and ‘‘value sets”, are the building blocks for creating the database queries and would mainly be used to: identify a cohort of patients for use in an observational or retrospective study; identify the covariates of interest, confounders and endpoints for the same observational study; or identify potentially at risk patients when used as part of a clinical quality measure (Akbarov et al., 2015).
Interoperability could be defined as the ability for computer systems (databases) to talk to each other and/or to different types of users such as physicians versus patients, adults versus children, and so on.
Syntactic interoperability: Does the data record the same thing between systems, for example Cerner codes gender: male = M, Female = F, Unknown = U Epic codes gender: Male = 0, Female = 1, Other = 3, Unknown =
Syntactically inoperable are on two levels: (a). Cerner does not account for “other”, and (b). Uses categorical features, not numeric features. This makes it impossible to directly merge these data sets into one system. It needs to be mapped to a data model to become interoperable (hence OMOP, HL7, FHIR etc.). Other obvious examples are height in feet (ft) and inches (in) vs. meters; weight in pounds (Ibs) and ounces (oz) vs. kilogram (kg).
Semantic interoperability: Does the data mean the same thing between systems: (a) Cerner = Discontinued_medication = patient has been discharged/died and hospital pharmacy no longer dispensing. (b) Epic = Discontinued_medication = patient is refusing medication, so order discontinued. This gives rise to semantic ontologies to force us to use a standard code set to communicate.
In addition, it has been ascertained that the size and complexity of the final set of the codes is determined by a combination of the particular code terminology and the concept of interest. For example, an evaluation of 1054 code sets created for clinical quality measures found varied sizes from sets containing a single code, to a code set for trauma which included 20,560 codes (Winnenburg & Bodenreider, 2013). The construction and validation of such lists is a non-trivial matter. The creation of sets of clinical codes for querying EHR datasets is a critical important phase of using pre-use data for research. It is usually the initial and a difficult step in research as errors are introduced at this stage of missing or wrongly specified codes could result in selection biases that progresses throughout subsequent analyses which may have a major impact in the research outcome (Nicholson, Tate, Koeling, & Cassell, 2011). For instance, rheumatoid arthritis code set differences could induce nearly a sevenfold difference in estimates of the incidence (Nicholson et al.,2011; Rodríguez, Tolosa, Ruigómez, Johansson, & Wallander, 2009).
National and international standards are everywhere in our society. Although in its beginning stages, healthcare has now also developed and met standards and interoperability guidelines of its own. Currently, there are many systems that fail to interoperate. This increases the risk of potential lethal errors in patient care. Government agencies on all levels have recognized this very problem and have put the man power and financial resources in place to overcome this extremely complex issue. While standards grounded in health information exchange may not be the only solution to National and Worldwide health care reform it certainly does assist in accomplishing the goal. Hence, collaboration between academia, industry, regulatory bodies, policy makers, patients, electronic health record vendors and hospitals is critical for a complete systems and data integration for meaningful use of electronic health records in clinical research to answer such questions as, “Is there any relationship between hospital characteristics and inpatient diabetic myocardial infarction outcome?”. This kind of research could describe the EHR extraction in institutions with disparate systems in different hospitals.
Significance
To the best of the authors’ knowledge, challenges involving EHR/EMR data merge have not been investigated among urban hospitals. This project will inform future research activities that encompass data merging or comparisons to highlight issues associated with system interoperability efforts in clinical research. Health services and policy research require access to complete, accurate, and timely patient and organizational data. Often times, the health-related datasets are created and held by diverse and disparate public, private organizations and individual researchers due to coding challenges. To overcome this barrier, the investigators believe that extracting meaningful information from separate multiple secondary datasets is needed. Though the authors had some challenges in matching and linking data from disparate systems, this paper provides effective clinical research outcomes between the DMV urban hospitals that would enable best practice sharing to decrease undesirable patient outcomes in future investigations. In addition, less is known about diabetic muscle infarction (DMI) than myocardial infarction (MI) generally. Data mining, advanced statistical modeling and predictive data analytics are essential in today’s emerging big data to knowledge (BD2K) research which were used to provide answers to the aims outlined below. Urban hospitals serve the medical needs of patients with diabetes, cardiovascular and other chronic diseases through a multidisciplinary approach to patient care. This study will also empower physicians and identify predictors of inpatient DMI outcomes by abstracting data on DMI patients and harmonizing them to a common data model. The datasets will help identify opportunities for targeted interventions for inpatients at risk for readmission, particularly in groups with disparate health outcomes. Furthermore, the results will provide a foundation for future studies exploring the best practices for systems and data interoperability to answer relevant clinical questions.
Innovation
Biomedical informatics is the use of data mobilized by nationwide adoption of EHR under the Affordable Care Act (ACA) which aligns well with the National Biomedical Research Act—S.2624-114 by Congress on BD2K and National Institute of Health and National Heart, Blood and Lung Institute (NIH NHBLI) objectives 3, 7 & 8. Biomedical informatics is the discipline that seeks to apply computer and communication technology to improve health. Special efforts are being made by some informatics researchers to integrate datasets from disparate systems with different racial and ethnic groups, and researchers from economically, socially, culturally or educationally disadvantaged backgrounds, into informatics careers and perform research that disproportionally affect the underserved (The Kaiser Family Foundation and the American College of Cardiology Foundation, 2002). This is the first research on trends of hospital characteristics and outcomes between DMV urban hospitals. However, using disparate systems was a challenge and was not undertaken due to lack of system integration or dataset interoperability. In addition, results from hospitals serving a large number in the DC area may provide an insight on patient outcomes to enable reduction in mortality rates in the diabetic population.
Method
Data Integration
Using Cerner Soarian from an institution with multiple hospitals, EMR data were extracted using an eXtensible Markup Language (XML) interpreter (parser) from XLM documents. A data parser (C, Java, SQL) was used to interpret and process the data. In addition, Natural Language processing (NLP) technologies were developed for data passage via XML to complete a modular project on diabetic myocardial infarction (DMI). However, the authors experienced challenges in the system integration of the urban hospitals as the re-use of EHR data for research is a relatively new field, and code set engineering is an issue that requires attention. This is reflected in the lack of interoperability and methodological literature on the subject despite its importance (Dave & Petersen, 2009). Recently, interoperability in this area has received more attention with Gulliford et al., (2009) calling for ‘‘greater transparency in the selection of lists of codes for different conditions, as well as sharing of code lists among researcher” (p. e7168). Unfortunately, several reviews have shown that code sets are rarely included in applied papers (Springate, 2014), In addition, the process by which the code sets have been constructed are not described well.
Data Cleaning
Normally, data abstracted from EHRs are notoriously “messy” and require extensive cleaning and cross-checking before they can be confirmed as useful and useable. The most common issue experienced was missing values. The first goal was to compile a metadata file on each of the three categories (diabetes, diabetic myocardial infarction, myocardial infarction). This metadata included information on missingness, as well as other information that were useful both in the analysis as stated in the objective, and to create a master data management file for sharing with the research community at the end of the study based on appropriate regulations. We have examined data from the urban hospitals disorder M Page (M page is one of the Health IT interventions available to bedside clinicians in the hospital; a browser record embedded in the Cerner interface; these are managed by the hospital’s clinical informatics staff and available to the investigators – both clinical and non-clinical). The M Page collects 83 variables for display for a maximum of two weeks of records (~150-200 SIRS and disorder alerts). A calendar-like function allows for historic data pulls if needed for validation. Demographics and vital data were complete in most cases.
Laboratory and vital sign data were somewhat variable ranging from 70 – 100% and MAP. The M page is limited in its display by the underlying logic of the commands pulling data from the Cerner database and we were confident that missing data in these cases could be pulled from ‘nearest neighbor’ records in the EHR. Data on lab orders was also variably missing, but again, other copies of orders exist in the record to fill gaps. Data generated from this application allowed further applications to fund advanced remedial training, as well as health IT and human factors research for bedside clinicians to emphasize the timely entry of observations to the EHR to (1) mitigate missingness in essential health IT systems and (2) enhance the functionality of Health IT systems by providing timely and accurate data, especially during the care of critically ill patients.
For serially missing data there were two options: impute or ignore. In serial observations on a close time scale, for example automated blood pressure or respiration measurements at the bedside, simple imputation techniques were used to infer missing data. We inferred missing values in these cases and visualized the serial values using Tableau software to ensure no excess deviation from biological possibility, as well as to ensure trends remained intact. All imputations and our observations of visualized data were recorded in the metadata file. Where it was not possible to impute missing data from closely linked variables we flagged data as missing. Although missing data are always a statistical problem when working with EHR data, we expected this research to gather a large enough volume of patient data that missingness would not skew our output.
A second common issue we faced was found on fields lacking proper automated front end validation. In these cases where there was no fixed range of values to ensure data were entered into the EHR correctly, transcription errors were common. In some cases, these were relatively easy to find, annotate and replace. An example was inverted systolic and diastolic blood pressures, or mixed heart and respiration rates. In other cases, data was annotated or flagged as potentially wrong, and managed statistically at the point of analysis. This included deleting variables that were clearly outside natural biological limits. Again, as with missing data we expected our research to gather a large enough volume of patient data that missingness did not skew our output.
Data harmonization
Cerner Millennium and GE Centricity EHR were merged in some hospitals as these two systems rely on different database structures and data models. We used the most current version of the Observational Health Data Sciences and Informatics Common Data Model (OHDSI-CDM; Version 3). The OHDSI CDM is a widely adopted and validated data model that uses standard medical and clinical vocabularies (ICD-10, CPT, LOINC, etc.) to ensure data were interoperable and confidential between sites and systems at a research institute. In many countries, researchers and administrators have struggled to apply a standardized data model to harmonize and collect medical data from a variety of heterogeneous sources. However, various barriers exist, such as system heterogeneity, various data formats, changes in human protection rules around the world, trust building, contracting and coordination, and research governance policies (Park, 2017).
The OHDSI CDM was recommended and required by NIH and the Patient-Centered Outcomes Research Institute (PCORI). This mapping served two purposes: first it allowed us to transform all of our data into one type allowing for easier analysis of the entire corpus, as opposed to being constrained to run repeated analyses against different data sets. Secondly, it made the data more broadly useful and useable by colleagues after our analysis was complete. Mapping to the OHDSI-CDM was carried out en masse using open source extract-transform-load tools; either Talend DI or Pentaho Kettle. These tools allow the desired end format to be programmed as an output type. The Cerner and legacy Centricity data files were mapped to the correct OHDSI-CDM terminology. The software then automatically transformed the data to the correct type and expressed an output file for analysis. We tested extensively on small data sets to ensure no information was lost during the translation process. For example, input of a decimal versus an integer led to loss of values due to rounding errors. Once the combined data sets were mapped to the same common data model they were stored as comma separated values (.csv) files and SAS format.
Limitations and Alternative Solutions
Although secondary data analysis requires extra rigour in data extraction, cleaning and analysis, the investigators were cognizant of the potential benefits that was derived from the proposed study. To overcome any technical and statistical challenges or research design barriers, the investigators were carefully selected and made up of an interdisciplinary team with expertise in biostatistics, biomedical informatics, and healthcare to work on the planning and conduct of this study. Another potential limitation emerged in the difficulty in accessing the institution(s) multiple health and operation systems and authenticating the hospitals’ accuracy in reporting coded procedures and processes. To address this, a good working relationship was established with key personnel at each of the institutions. Data interoperability of some systems was ensured through the use of standard medical vocabularies such as ICD-10-CM diagnostic coding, CPT procedure coding and drug libraries (LOINC, NDC etc.). Extracted data were mapped to a common data model using open source extract/transform/load tools. All investigators involved in the project were mandated to undergo research integrity (basic biomedical certificates), and privacy and confidentiality (HIPAA) training. Data collected during this project were stored in encrypted databases behind institutional firewalls and accessed using password protected web-services. All hardware (laptops) were physically secured and also featured password protection and full disk encryption to ensure privacy and confidentiality.
Study Variable Measures
Patient Measures: Age 20-80 years. Gender (dichotomous): Male, Female. Race (categorical): white, black, other. Other measures included Body mass index (BMI), Systolic blood pressure (SBP), Diastolic blood pressure (DBP), Low density lipoprotein (LDL), High density lipoprotein (HDL), Hospitals (categorical), and Hospital categories (A & B).
Outcomes Measures: Mortality was defined as patient dying in the hospital.
Statistical Analysis
Data was extracted for disease categories to enable comparative outcomes studies within and in between diseases groups. The rationale for this was that there has been controversial discussion on the severity outcomes of DM only, MI only and both with the assumptions that MI category may have more detrimental outcomes than both DM and DMI. Descriptive statistics (frequencies, percentages, mean with standard deviation) as appropriate were used to summarize the data. ANOVA was used to compare differences in means of specific clinical occurrences between the three disease categories- DM, MI, DM+MI. Mortality rates due to DM, MI and DM + MI were evaluated and compared using a Chi-Square test. In addition, we performed logistic regression analyses to examine the independence of disease category in predicting mortality in models that adjusted for age, gender, and race. All of the analyses were conducted using IBM SPSS software (version 25.0, IBM SPSS). All tests were two-tailed at the 5% (p=0.05) level of significance.
Results
Of the 4,350 patients admitted with diabetes and myocardial infarction in the urban hospitals, 2,680 were diagnosed with diabetes (DM) only, 618 with myocardial infarction (MI) only, while 1,052 had both diabetes and Myocardial infarction (DM-MI). The demographic distribution showed that males (59.4%) were admitted more for the diabetes category than females (40.6%). In contrast, females were admitted more for MI only and DM + MI. It is also important to note that younger individuals tended to be in the diabetic category among hospital category A inpatients when compared to category B. Black Americans were 300 times more likely compared to white counterparts to be in the diabetes category. However, there was no significant difference between males and females for MI only and DM + MI (Table 1).
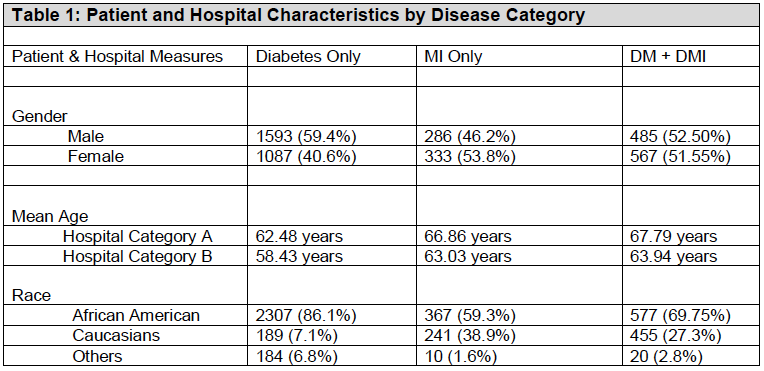
Furthermore, Table 2 shows that age, gender and race distribution were statistically significant in the DM only category with a p-value (<0.001) when compared to the MI only category for age (p=0.005) and DM + MI category for age (p=0.027) and race (p<0.000). For the clinical measures, DBP and HDP were statistically significant at p <0.001 for DM only. For MI only, the statistical significance ranged from p <0.001 to 0.023 with SBP having the highest significance while LDL was not statistically significant (p-0.738). DM + MI showed the highest statistical p-value (0.025) for BMI.
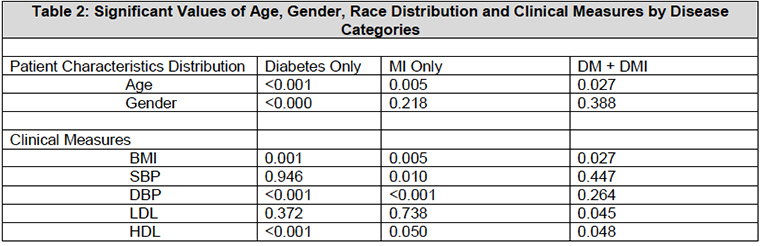
BMI—Body Mass Index
SBP—Systolic Blood Pressure
DBP—Diastolic Blood Pressure
LDL—Low Density lipoprotein
HDL—High Density lipoprotein
Table 3 shows that hospital category A and B comparative analysis resulted in p-values of (0.000) in age, BMI, DBP and HDL. However, there were no statistical difference noted for both SBP and LDL clinical measures.
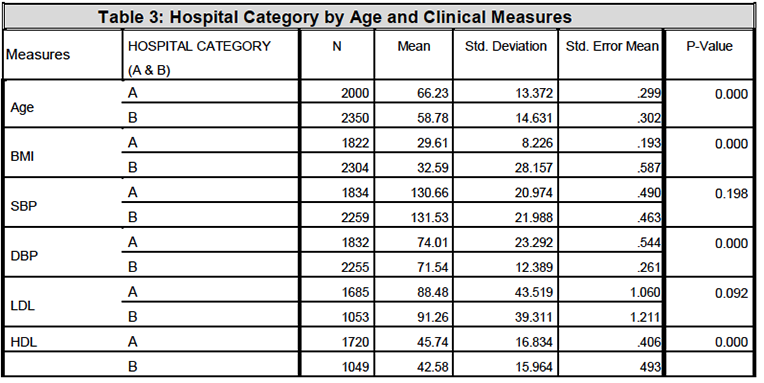
The odds ratio analysis showed that there is a statistical significance for lower risk of death from the DM category in hospital category A when compared to other disease categories. However, it is important to note that the risk of death for MI only was not statistically significant for both hospital category A and B. Furthermore, there was no risk of death for DM + MI in hospital category A (Table 4-7, see the Mantel – Haenzel Common odds Ratio).
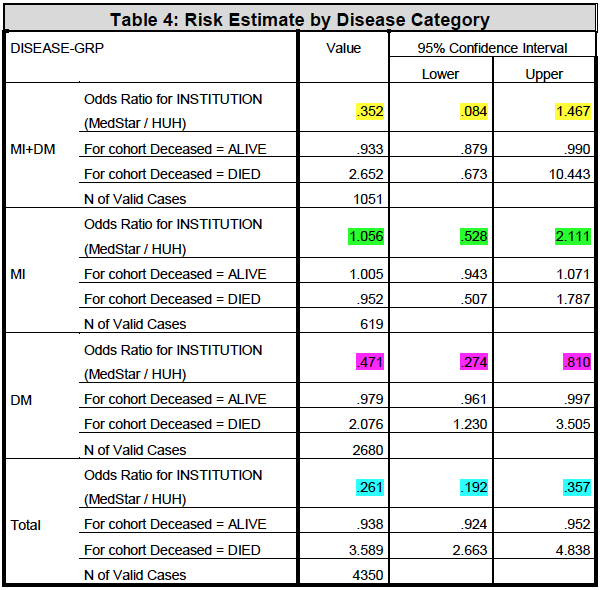
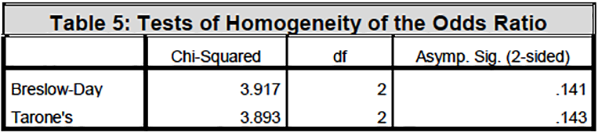
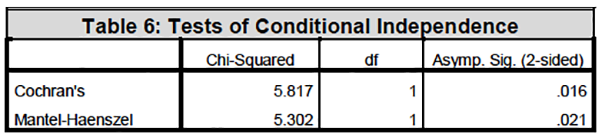
Under the conditional independence assumption, Cochran’s statistic is asymptotically distributed as a 1 df chi-squared distribution, only if the number of strata is fixed, while the Mantel-Haenszel statistic is always asymptotically distributed as a 1 df chi-squared distribution.
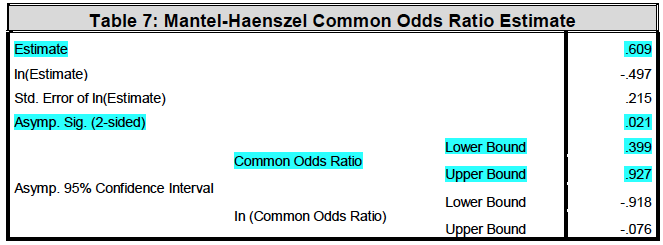

Discussion & Conclusion
Though the advancement of EHR utilization for clinical research may have opened the door for study feasibility and the streamlining of data collection at baseline and follow-up points, it is worth noting that that there have been concerns regarding some challenges such as burdensome systems interoperability, obstructive data collection, and uncertain generalizability of results. However, it is also important to note that leveraging EHRs to counterbalance these trends is an area of intense interest and data sharing from urban hospitals may enable others to learn best practices between and among hospitals. To inform this issue, we have examined differences in data mining of urban hospital categories (A & B) for 2,013 discharged patients with diabetes and myocardial infarction. The results of this study showed that some of these hospital categories may have healthcare practices that others may choose to follow to improve inpatient outcomes and disease outcomes. Hence, best practices and procedures could be shared among urban hospitals in order to optimize the use of EHR data in pilot and population studies.
Acknowledgement
This project has been funded in whole or in part with Federal funds (UL1TR000101 previously UL1RR031975) from the National Center for Advancing Translational Sciences (NCATS), National Institutesof Health, through the Clinical and Translational Science Awards Program (CTSA), a trademark of DHHS, part of the Roadmap Initiative, “Re-Engineering the Clinical ResearchEnterprise.
References
Akbarov, A., Kontopantelis, E., Sperrin, M., Stocks, S.J., Williams, R., Rodgers, S., Avery, A., Buchan, I. & Ashcroft, D.M. (2015). Primary care medication safety surveillance with integrated primary and secondary care electronic health records: a cross-sectional study, Drug Safety, 38, 671–682, s40264- 015-0304-x.
Dave, S. & Petersen, I. (2009). Creating medical and drug code lists to identify cases in primary care databases, Pharmacoepidemiology and drug safety,18, 704–707,http://dx.doi.org/10.1002/pds.1770.
Gulliford. M..C., Charlton, J., Ashworth, M., Rudd, A.G., Toschke, A.M., Delaney, B., Grieve, A, Heuschmann, P.U., Little, P., Redfern, J., van Staa, T., Wolfe, C., Yardley, L &, McDermott, L (2009), Selection of medical diagnostic codes for analysis of electronic patient records. Application to stroke in a primary care database, PLoS ONE 4(9), e7168. ,http://dx.doi.org/10.1371/journal.pone.0007168.
National Institute of Health and National Heart, Blood and Lung Institute (NIH NHBLI). (2019).Big Data to Knowledge. https://commonfund.nih.gov/bd2k
Nicholson, A., Tate, A.R., Koeling, R. & Cassell, J.A (2011). What does validation of cases inelectronic record databases mean? The potential contribution of free text, Pharmacoepidemiology and drug safety. 20, 321–324, http://dx.doi.org/10.1002/pds.2086.
Park, R.W. (2017). Sharing Clinical Big Data While Protecting Confidentiality and Security: Observational Health Data Sciences and Informatics. Healthcare Informatics Research, 23(1):1-3.
Rodríguez, L.A.G., Tolosa, L.B., Ruigómez, A., Johansson, S. & Wallander, M.A. (2009). Rheumatoidarthritis in UK primary care: incidence and prior morbidity, Scandinavian Journal of Rheumatology, 38,173–177.http://dx.doi.org/10.1080/03009740802448825.
Springate, D.A., Kontopantelis, E., Ashcroft, D.M., Olier, I., Parisi, R., Chamapiwa, E. & Reeves, D. (2014). ClinicalCodes: an online clinical codes repository to improve the validity and reproducibility of research using electronic medical records, PLoS ONE, 9(6), e99825,http://dx.doi.org/10.1371/journal.pone.0099825.
TheKaiserFamilyFoundationandtheAmericanCollegeofCardiologyFoundation.(2002). Racial/Ethnic Differences in Cardiac Care: The Weight of the Evidence.(Report #6040) https://www.kff.org/disparities-policy/fact-sheet/racialethnic-differences-in-cardiac-care-the-weight/
Williams, R., Kontopantelis, E., Buchan, I., & Peek, N. (2017). Clinical code set engineering for reusing EHR data for research: A review. Journal of Biomedical Informatics, 70, 1–13.
Winnenburg, R. & Bodenreider, O (2013). Metrics for assessing the quality of value sets in clinical quality measures. AMIA Annual Symposium proceedings, 1497–1505.






